Analysis software processes the raw data and assigns bases to the peaks to generate the final trace and sequence. Signal intensity, in relative fluorescence units, is plotted over time, which correlates with base position. Fluorescence is detected at the end of the capillary, and signal intensity from four color channels, each representing a DNA base, is plotted on the y-axis relative to time on the x-axis.Īnatomy of a chromatogram. Here, we provide a guide to understanding Sanger sequencing data, covering the following topics:Ī chromatogram represents the migration of labeled sequencing products via capillary electrophoresis. Thus, it's important to visually inspect all your traces to ensure that the output sequence represents reality. The base-calling software does its best to interpret the chromatogram, but it's not always accurate, especially with poor-quality data. The chromatogram contains valuable data that speaks to the accuracy of the generated sequence. Although the latter may seem to hold all the relevant information-after all, the point of sequencing is to get a sequence-the former can't be ignored. The output for Sanger sequencing is typically a chromatogram, also known as a trace or ab1 file, and a text-based sequence file. Business Continuity and Risk Mitigation.Sample and Material Management Solutions.Sample Transport and Cold-Chain Logistics.Discovery, Compound Management and Biologics.The Java language version of PatternHunter is just 40 KB, only 1% the size of Blast, while offering a large portion of its functionality. Its features are its advanced patented algorithm and data structures, and the java language used to create it. PatternHunter PatternHunter, based on Java, can identify all approximate repeats in a complete genome in a short time using little memory on a desktop computer. PROSPECT PROSPECT (PROtein Structure Prediction and Evaluation Computer ToolKit) is a protein-structure prediction system that employs a computational technique called protein threading to construct a protein's 3-D model. Protein Explorer, a derivative of RasMol, is an easier to use program. 
RasMol It is a powerful research tool to display the structure of DNA, proteins, and smaller molecules. 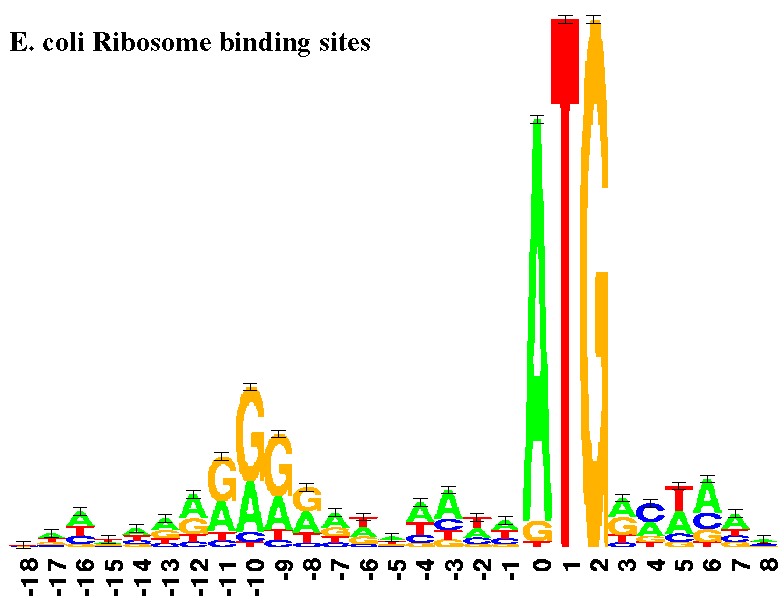
It returns the best match over a total length of input sequences, be it a protein or a nucleic acid. Clustalw It is a fully automated sequence alignment tool for DNA and protein sequences. It provides a set of sequence-analysis programs, and also supports all UNIX platforms. Extensive libraries are also provided with this package, allowing other scientists to release their software as open source. It can work with data in a range of formats and also retrieve sequence data transparently from the Web. EMBOSS EMBOSS (European Molecular Biology Open Software Suite) is a software-analysis package. NCBI has also introduced the new queuing system to BLAST (Q BLAST) that allows users to retrieve results at their convenience and format their results multiple times with different formatting options. Also a protein database can be searched to find a match against the queried protein sequence. Comparison of nucleotide sequences in a database can be performed. Perform fast similarity searches regardless of whether the query is for protein or DNA. It is a set of search programs designed for the Windows platform and is used to BLAST ( Basic Local Alignment Search Tool) BLAST ( Basic Local Alignment Search Tool) comes under the category of homology and similarity tools. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
